But the new study shows that different P450s selectively partner with different electron donors (and electron transport chains) to drive their activities. These electrons act as a power source to fuel the machine," Liu explained.Ĭonventionally, scientists thought that the P450s primarily interact with a general electron donor called cytochrome P450 reductase to produce a variety of aromatic compounds. "To make P450 machines run, they need partner molecules to deliver electrons. Scientists have long known that P450 enzymes do not work alone to determine the structural and biological features of aromatic compounds. 
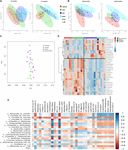
The work could help facilitate long-term carbon storage and the carbon-neutral utilization of plant biomass for energy applications, improve plants' nutritional properties, or increase their resistance to disease and harsh environmental conditions. Uncovering the complexity of how these enzymes are regulated provides a new set of genetic tools scientists can use to precisely control which compounds get produced in different parts of a plant. "These enzymes operate as a synthetic machine to produce a wide range of aromatic compounds in plants-including compounds that build plants' waterproof skeleton and vasculature, and others that provide defense from insect invasions and ultraviolet (UV) radiation." "Our study reveals the long-overlooked complexity and versatility of a key set of enzymes known as cytochrome P450 monooxygenases," said study lead author Chang-Jun Liu of Brookhaven Lab's biology department. The research, just published in the journal Science Advances, suggests new strategies for controlling plant biochemistry for agricultural and industrial applications. Department of Energy's Brookhaven National Laboratory have discovered a new level of regulation in the biochemical "machinery" that plants use to convert organic carbon derived from photosynthesis into a range of ring-shaped aromatic molecules. (In both images the red signal comes from chlorophyl.). In this case the scientists attached half of the GFP tag to each of these proteins the fluorescent glow occurs only when the two halves come together as the proteins interact. Right: A GFP-labeled complex of the P450 enzyme interacting with the electron donor. Left: Localization of a GFP-labeled electron donor protein along the endoplasmic reticulum (inner network of membranes) in leaf cells. Given the distinctive bacterial species and strain composition found in each individual's gut, our findings suggest the IgA antibody repertoire is shaped uniquely to bind "self" gut bacteria.In addition to performing genetic and biochemical studies, the scientists used a green fluorescent protein (GFP) "tag" and microscope facilities at Brookhaven Lab's Center for Functional Nanomaterials to visualize the proteins they were studying in leaf cells. Species-specific IgAs had a range of strain specificities. Orally administered hybridoma-produced IgAs still retained bacterial antigen binding capability, implying the potential for a new class of therapeutic antibodies. IgA hybridomas generated from lamina propria B cells of gnotobiotic mice showed that most IgA clones recognized a single bacterial species, whereas a small portion displayed cross-reactivity. By colonizing germ-free mice with defined commensal bacteria, we found that the binding specificity of bulk fecal and serum IgA toward resident gut bacteria resolves well at the species level and has modest strain-level specificity. Despite being the most abundantly secreted immunoglobulin isotype, the pattern of reactivity of immunoglobulin A (IgA) antibodies toward each individual's own gut commensal bacteria still remains elusive.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
